MMAPPR2: Mutation Mapping from RNA Sequences
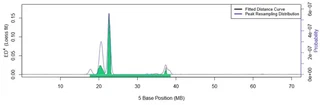 Localizing a mutation on zebrafish chromosome 5 using genotype Euclidean distance (ED) between mutant and wild-type strains, processed with smoothing and resampling.
Localizing a mutation on zebrafish chromosome 5 using genotype Euclidean distance (ED) between mutant and wild-type strains, processed with smoothing and resampling.MMAPPR2 maps mutations resulting from pooled RNA-seq data from the F2 cross of forward genetic screens. It accepts aligned BAM files as well as a reference genome as input and identifies loci of high sequence disparity between the control and mutant RNA sequences. It is significantly faster than its predecessor (Hill et al. 2013) and enhances its functionality by improving mutation localization and integrating with Ensembl’s Variant Effect Predictor, outputting a ranked list of candidate mutations.

Biomedical Engineering
I’m a PhD student advised by Chris Rozell at Georgia Tech, developing advanced closed-loop optogenetic control techniques for neuroscience. Specifically, I am applying optimal, model-based control to precisely perturb population dynamics.
I’m fascinated by the biological computational principles and how we can exploit them for next-generation AI/ML technologies. And vice-versa: how we can apply modern ML to neurotech/neuroscience, especially for building life- and time-saving brain-computer interfaces. I am looking for an industry research scientist role post-graduation.